2026
90
How a coral holobiont stays in sync in nature
Chan CX
2026. Cell Host & Microbe. 34: 190-191 • DOI • PDF
89
RNAcentral in 2026: Genes and literature integration
RNAcentral Consortium: Green A, Ribas CE, Jandalala I, Muston P, O’Cathail C, Cochrane G, Ernst C, Zhao L, Madrigal P, Attrill H, Marygold S, Lancet D, Dobzinski N, Chan PP, Lowe TM, Bruford EA, Seal RL, Hermjakob H, Panneerselvam K, Finn RD, Gurbich TA, Griffiths-Jones A, Fromm B, Peterson KJ, Sordyl D, Bujnicki JM, Velankar S, Appasamy SD, Ganguly S, Zhang P, He S, Rutherford KM, Wood V, Lovering RC, Picardi E, Ontiveros N, Huang L, Miao Z, Petrov AS, McCann H, Cavalleri E, Mesiti M, Rivas E, Szikszai M, Magnus M, Gerken J, Chuvochina M, Bergeron D, Scott M, Williams K, Gutell RR, Chan CX, Quinton-Tulloch M, Diamantakis S, Petrov AI, Bateman A, Sweeney BA
2026. Nucleic Acids Research. 54 (D1): D303-D313 • DOI • PubMed • PDF
2025
88
Insights into phylogeny, diversity and functional potential of Poseidoniales viruses
Prabhu A, Zaugg J, Chan CX, McIlroy SJ, Rink C
2025. Environmental Microbiology. 27: e70017 • DOI • PubMed • PDF
2024
87
Progress and future directions for seaweed holobiont research
Saha M, Dittami SM, Chan CX, Raina JB, Stock W, Ghaderiardakani F, VB AM, Corr S, Schleyer G, Todd J, Cardini U, Bengtsson MM, Prado S, Skillings D, Sonnenschein EC, Engelen AH, Wang G, Wichard T, Brodie J, Leblanc C, Egan S
2024. New Phytologist. 244: 364-376 • DOI • PubMed • PDF
86
Chromosomal inversions harbour excess mutational load in the coral, Acropora kenti, on the Great Barrier Reef
Zhang J, Schneller NM, Field MA, Chan CX, Miller DJ, Strugnell JM, Riginos C, Bay L, Cooke IR
2024. Molecular Ecology. 33: e17468. DOI • PubMed • PDF
85
Whole-genome duplication in an algal symbiont bolsters coral heat tolerance
Dougan KE, Bellantuono AJ, Kahlke T, Abbriano RM, Chen Y, Shah S, Granados-Cifuentes C, van Oppen MJH, Bhattacharya D, Suggett DJ, Rodriguez-Lanetty M, Chan CX
2024. Science Advances. 10: adn2218. • DOI • PubMed • PDF
84
Massive genome reduction predates the divergence of Symbiodiniaceae dinoflagellates
Shah S, Dougan KE, Chen Y, Lo R, Laird G, Fortuin MDA, Rai SK, Murigneux V, Bellantuono AJ, Rodriguez-Lanetty M, Bhattacharya D, Chan CX
2024. The ISME Journal. 18: wrae059. • DOI • PubMed • PDF
83
Nuclear genomes of dinoflagellates reveal evolutionarily conserved pattern of RNA editing relative to stress response
Chen Y, Dougan KE, Bhattacharya D, Chan CX
2024. Frontiers in Protistology. 2: 1320917. • DOI • PDF
82
Contaminant or goldmine? In silico assessment of Symbiodiniaceae community using coral hologenomes
Ishida H, Riginos C, Chan CX
2024. Frontiers in Protistology. 2: 1376877. DOI • PDF
81
Genome-wide transcriptome analysis reveals the diversity and function of long non-coding RNAs in dinoflagellates
Chen Y, Dougan KE, Nguyen Q, Bhattacharya D, Chan CX
2024. NAR Genomics and Bioinformatics. 6: lqae016. DOI • PubMed • PDF
80
Facultative lifestyle drives diversity of coral algal symbionts
Bhattacharya D, Stephens TG, Chille EE, Benites LF, Chan CX
2024. Trends in Ecology and Evolution. 39: 239-247. DOI • PubMed • PDF
2023
79
Multi-omics analysis reveals the molecular response to heat stress in a “red tide” dinoflagellate
Dougan KE*, Deng ZL*, Wöhlbrand L*, Reuse C*, Bunk B, Chen Y, Hartlich J, Hiller K, John U, Kalvelage J, Mansky J, Neumann-Schaal M, Overmann J, Petersen J, Sanchez-Garcia S, Schmidt-Hohagen K, Shah S, Spröer C, Sztajer H, Wang H, Bhattacharya D, Rabus R, Jahn D, Chan CX^, Wagner-Döbler I^
*: these authors contribute equally; ^: senior corresponding authors
2023. Genome Biology. 24: 265. DOI • PDF
78
Developing model systems for dinoflagellates in the post-genomic era
Ishida H, John U, Murray MA, Bhattacharya D, Chan CX
2023. Journal of Phycology. 59: 799-808. DOI • PubMed • PDF
77
Gene duplication is the primary driver of intraspecific genomic divergence in coral algal symbionts
Shah S, Dougan KE, Chen Y, Bhattacharya D, Chan CX
2023. Open Biology. 13: 230182. DOI • PubMed • PDF
76
In vivo structure probing of RNA in Archaea: novel insights into the ribosome structure of Methanosarcina acetivorans
Williams AM, Jolley E, Santiago-Martínez MG, Chan CX, Gutell R, Ferry JG, Bevilacqua PC
2023. RNA. 29: 1610-1620. DOI • PubMed • PDF
75
Building consensus around the assessment and interpretation of Symbiodiniaceae diversity
Davies SW, Gamache MH, Howe-Kerr LI, Kriefall NG, Baker AC, Banaszak AT, Bay LK, Bellantuono AJ, Bhattacharya D, Chan CX, Claar DC, Coffroth MA, Cunning R, Davy SK, del Campo J, Díaz-Almeyda EM, Frommlet JC, Fuess LE, González-Pech RA, Goulet TL, Hoadley KD, Howells EJ, Hume BCC, Kemp DW, Kenkel CD, Kitchen SA, LaJeunesse TC, Lin S, McIlroy SE, McMinds R, Nitschke MR, Oakley CA, Peixoto RS, Prada C, Putnam HM, Quigley KM, Reich HG, Reimer JD, Rodriguez-Lanetty M, Rosales SM, Saad OS, Sampayo EM, Santos SR, Shoguchi E, Smith EG, Stat M, Stephens TG, Strader ME, Suggett DJ, Swain TD, Tran C, Traylor-Knowles N, Voolstra CR, Warner ME, Weis VM, Wright RM, Xiang T, Yamashita H, Ziegler M, Correa AMS, Parkinson JE
2023. PeerJ 11: e15023. DOI • PubMed • PDF
74
Micro‑analytical and molecular approaches for understanding the distribution, biochemistry, and molecular biology of selenium in (hyperaccumulator) plants
Pinto Irish K, Harvey MA, Harris HH, Aarts MGM, Chan CX, Erskine PD, van der Ent A
2023. Planta 257: 2. DOI • PubMed • PDF
2022
73
Evolutionary responses of a reef-building coral to climate change at the end of the last glacial maximum
Zhang J, Richards ZT, Adam AAS, Chan CX, Shinzato C, Gilmour J, Thomas L, Strugnell JM, Miller DJ, Cooke I
2022. Molecular Biology and Evolution 39: msac201. DOI • PubMed • PDF
72
Improved Cladocopium goreaui genome assembly reveals features of a facultative coral symbiont and the complex evolutionary history of dinoflagellate genes
Chen Y, Shah S, Dougan KE, van Oppen MJH, Bhattacharya D, Chan CX
2022. Microorganisms 10: 1662. • DOI • PubMed • PDF
71
Genome-powered classification of microbial eukaryotes: focus on coral algal symbionts
Dougan KE, González-Pech RA, Stephens TG, Shah S, Chen Y, Ragan MA, Bhattacharya D, Chan CX
2022. Trends in Microbiology • 30: 831-840 • DOI • PubMed • PDF (accepted manuscript)
70
Transcriptome of the coralline alga Calliarthron tuberculosum (Corallinales, Rhodophyta) reveals convergent evolution of a partial lignin biosynthesis pathway
Xue JY, Hind K, Lemay MA, Mcminigal A, Jourdain E, Chan CX, Martone PT
2022. PLoS ONE 17: e0266892. • DOI • PubMed • PDF
69
Genome-guided analysis of seven weed species reveals conserved sequence and structural features of key gene targets for herbicide development
Shah S, Lonhienne T, Murray CE, Chen Y, Dougan KE, Low YS, Williams CM, Schenk G, Walter GH, Guddat LW, Chan CX
2022. Frontiers in Plant Science • 13: 909073. • DOI • PubMed • PDF
68
Alignment-free analysis of whole-genome sequences from Symbiodiniaceae reveals differential phylogenetic signals in distinct regions
Lo R, Dougan KE, Chen Y, Shah S, Bhattacharya D, Chan CX
2022. Frontiers in Plant Science • 13: 815714 • DOI • PubMed • PDF
67
Nuclear genome of a pedinophyte pinpoints genomic innovation and streamlining in the green algae
Repetti S, Iha C, Uthanumallian K, Jackson C, Chen Y, Chan CX, Verbruggen H
2022. New Phytologist • 233: 2144–2154. • DOI • PubMed • PDF
66
Tightly constrained genome reduction and relaxation of purifying selection during secondary plastid endosymbiosis
Uthanumallian K, Iha C, Repetti SI, Chan CX, Bhattacharya D, Duchene S, Verbruggen H
2022. Molecular Biology and Evolution • 39(1): msab295. • DOI • PubMed • PDF
2021
65
Consensus guidelines for advancing coral holobiont genome and specimen voucher deposition
Voolstra CR, Quigley KM, Davies SW, Parkinson JE, Peixoto RS, Aranda M, Baker AC, Barno AR, Barshis DJ, Benzoni F, Bonito V, Bourne DG, Buitrago-López C, Bridge TCL, Chan CX, Combosch DJ, Craggs J, Frommlet JC, Herrera S, Quattrini AM, Röthig T, Reimer JD, Rubio-Portillo E, Suggett DJ, Villela H, Ziegler M, Sweet M
2021. Frontiers in Marine Science • 8: 701784 • DOI • PDF
64
Comparative genomics supports that Brazilian bioethanol Saccharomyces cerevisiae comprise a unified group of domesticated strains related to cachaça spirit yeasts
Jacobus AP, Stephens TG, Youssef P, González-Pech RA, Ciccotosto-Camp MM, Dougan KE, Chen Y, Basso LC, Frazzon J, Chan CX, Gross J
2021. Frontiers in Microbiology • 12: 644089. • DOI • PubMed • PDF
63
Comparison of 15 dinoflagellate genomes reveals extensive sequence and structural divergence in family Symbiodiniaceae and genus Symbiodinium
González-Pech RA, Stephens TG, Chen Y, Mohamed AR, Cheng Y, Shah S, Dougan KE, Fortuin MDA, Lagorce R, Burt DW, Bhattacharya D, Ragan MA, Chan CX
2021. BMC Biology • 19: 73. • DOI • PubMed • PDF
62
Genomic adaptations to an endolithic lifestyle in the coral-associated alga Ostreobium
Iha C, Dougan KE, Varela JA, Avila V, Jackson CJ, Bogaert KA, Chen Y, Judd LM, Wick R, Holt KE, Pasella MM, Ricci F, Repetti SI, Medina M, Marcelino VR, Chan CX, Verbruggen H
2021. Current Biology • 31(7): 1393-1402 • DOI • PubMed
2020
61
Dual RNA-seq analyses of a coral and its native symbiont during the establishment of symbiosis
Mohamed AR, Andrade N, Moya A, Chan CX, Negri A, Bourne DG, Ying H, Ball EE, Miller DJ
2020. Molecular Ecology • 29: 3921-3937. • DOI • PubMed
60
Comparative transcriptomic analyses of Chromera and Symbiodiniaceae
Mohamed AR, Chan CX, Ragan MA, Zhang J, Cooke I, Ball EE, Miller DJ
2020. Environmental Microbiology Reports • 12: 435-443. • DOI • PubMed • PDF
59
Sex in Symbiodiniaceae dinoflagellates: genomic evidence for independent loss of the canonical synaptonemal complex
Shah S, Chen Y, Bhattacharya D, Chan CX
2020. Scientific Reports • 10: 9792. DOI • PubMed • PDF
Featured in:
IMB News: Algae use distinct method when swapping DNA during sex
58
Genomes of the dinoflagellate Polarella glacialis encode tandemly repeated single-exon genes with adaptive functions
Stephens TG, González-Pech RA, Cheng Y, Mohamed AR, Burt DW, Bhattacharya D, Ragan MA, Chan CX
2020. BMC Biology • 18: 56. DOI • PubMed • PDF
Featured in:
IMB News: Algae use distinct method when swapping DNA during sex
57
Evidence that inconsistent gene prediction can mislead analysis of dinoflagellate genomes
Chen Y, González-Pech RA, Stephens TG, Bhattacharya D, Chan CX
2020. Journal of Phycology • 56: 6-10. DOI • PubMed • PDF • Data Site
2019
56
A genomic view of the reef-building coral Porites lutea and its microbial symbionts
Robbins SJ, Singleton CM, Chan CX, Messer LF, Geers AU, Ying H, Baker A, Bell SC, Morrow KM, Ragan MA, Miller DJ, Forêt S, ReFuGe2020 Consortium, Voolstra CR, Tyson GW, Bourne DG
2019. Nature Microbiology • 4: 2090-2100. • DOI • PubMed • PDF • Data Site
Featured in:
UQ News: Scientists decode DNA of coral and all its microscopic supporters
IMB News: Genetics of coral a key to saving reefs from climate change
Nature Microbiology Community – Behind the Paper: A genomic view of a coral holobiont
Cosmos Magazine: Corals need a lot of help from their friends
United Press International: DNA analysis details relationships between coral, algae and bacteria
Sci-News: Researchers sequence genomes of reef-building coral and its microbial symbionts
55
Benchmarking of alignment-free sequence comparison methods
Zielezinski A, Girgis HZ, Bernard G, Leimeister CA, Tang K, Dencker T, Lau AK, Röhling S, Choi J, Waterman MS, Comin M, Kim SH, Vinga S, Almeida JS, Chan CX, James BT, Sun F, Morgenstern B, Karlowski WM
2019. Genome Biology • 20: 144. • DOI • PubMed • PDF • AFproject site
54
Analysis of an improved Cyanophora paradoxa genome assembly
Price DC, Goodenough UW, Roth R, Lee JH, Kariyawasam T, Mutwil M, Ferrari C, Facchinelli F, Ball SG, Cenci U, Chan CX, Wagner NE, Yoon HS, Weber APM, Bhattacharya D
2019. DNA Research • 26: 287-299. • DOI • PubMed • PDF
53
Genome evolution of coral reef symbionts as intracellular residents
González-Pech RA, Bhattacharya D, Ragan MA, ChanCX
2019. Trends in Ecology & Evolution • 34: 799-806 • DOI • PubMed • Publisher • PDF (accepted manuscript)

Featured in:
UQ News: Understanding relationship break-ups to protect the reef
IMB News: Coral and algae relationship the key to understanding bleaching
United Press: Better understanding of coral-algae relationship could help prevent bleaching
Brisbane Times: Good relationships under the sea could hold key to coral bleaching
Sydney Morning Herald: Good relationships under the sea could hold key to coral bleaching
Space Daily: Better understanding of coral-algae relationship could help prevent bleaching
52
Molecular techniques and their limitations shape our view of the holobiont
Cooke I, Mead O, Whalen C, Boote C, Moya A, Ying H, Robbins S, Strugnell JM, Darling A, Miller DJ, Voolstra CR, Adamska M, Consortium of Australian Academy of Science Boden Research Conference Participants
2019. Zoology • 137: 125695 • DOI • PubMed • PDF
51
Resolving structure and function of metaorganisms through a holistic framework combining reductionist and integrative approaches
Jaspers C, Fraune S, Arnold AE, Miller DJ, Bosch TCG, Voolstra CR, Consortium of Australian Academy of Science Boden Research Conference Participants
2019. Zoology • 133: 81-87 • DOI • PubMed • PDF
50
Commonly misunderstood parameters of NCBI BLAST and important considerations for users
González-Pech RA, Stephens TG, Chan CX
2019. Bioinformatics • 35: 2697-2698. DOI • PubMed • PDF (accepted manuscript)
49
Alignment-free inference of hierarchical and reticulate phylogenomic relationships
Bernard G, Chan CX, Chan YB, Chua XY, Cong Y, Hogan JM, Maetschke SR, Ragan MA
2019. Briefings in Bioinformatics • 20: 426-435. • DOI • PubMed • PDF
2018
48
Core genes in diverse dinoflagellate lineages include a wealth of conserved dark genes with unknown functions
Stephens TG, Ragan MA, Bhattacharya D, Chan CX
Scientific Reports • 8: 17175. • DOI • PubMed • PDF
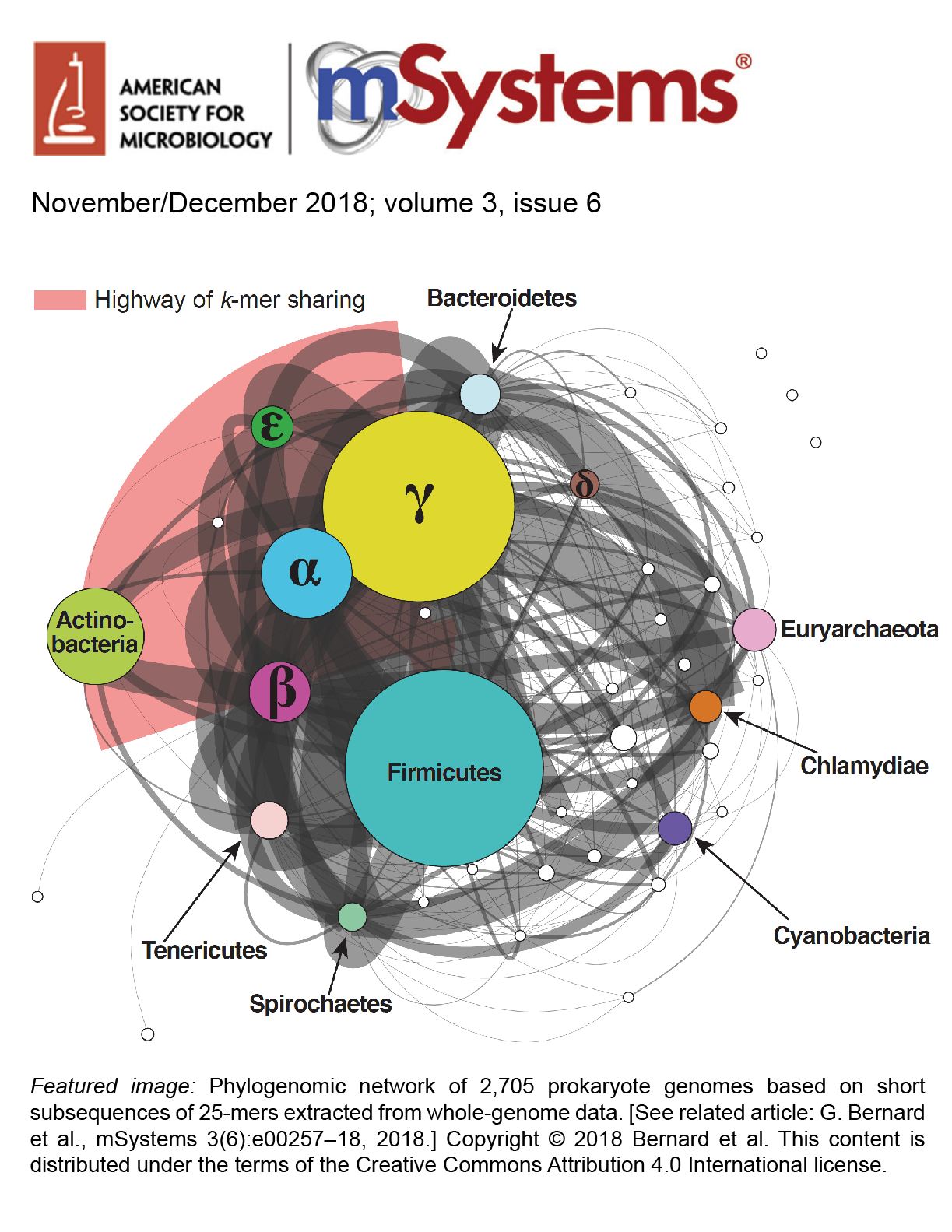
47
k-mer similarity, networks of microbial genomes and taxonomic rank
Bernard G, Greenfield P, Ragan MA, Chan CX
mSystems • 3: e00257-18. • DOI • PubMed • PDF • Supplemental Site
46
Symbiodinium genomes reveal adaptive evolution of functions related to coral-dinoflagellate symbiosis
Liu H, Stephens TG, González-Pech RA, Beltran VH, Lapeyre B, Bongaerts P, Cooke I, Aranda M, Bourne DG, Forêt S, Miller DJ, van Oppen MJH, Voolstra CR, Ragan MA, Chan CX
2018. Communications Biology • 1: 95. • DOI • PubMed • PDF • Behind-the-Paper •
Featured in:
IMB News: Unlocking genetic secrets to help save the Great Barrier Reef
GBRF: Decoding the Reef leads to world first—Unlocking genetic secrets of coral symbiosis to preserve iconic Great Barrier Reef
Video Feature
Featured in IMB 2018 year in review
45
Analysis of the draft genome of the red seaweed Gracilariopsis chorda provides insights into genome size evolution in Rhodophyta
Lee J, Yang EC, Graf L, Yang JH, Qiu H, Zelzion U, Chan CX, Stephens TG, Weber APM, Boo GH, Boo SM, Kim KM, Shin Y, Jung M, Lee SJ, Yim HS, Lee JY, Bhattacharya D, Yoon HS
2018. Molecular Biology and Evolution • 35: 1869-1886. • DOI • PubMed • PDF
44
Active host response to algal symbionts in the sea slug Elysia chlorotica
Chan CX, Vaysberg P, Price DC, Pelletreau KN, Rumpho ME, Bhattacharya D
2018. Molecular Biology and Evolution • 35: 1706-1711. • DOI • PubMed • Full text • PDF • National Geographic Feature • Video Feature
43
Plastid phylogenomics with broad taxon sampling further elucidates the distinct evolutionary origins and timing of secondary green plastids
Jackson C, Knoll AH, Chan CX, Verbruggen H
2018. Scientific Reports • 8: 1523. • DOI • PubMed • PDF
42
Deciphering the nature of the coral-Chromera association
Mohamed AR, Cumbo V, Harii S, Shinzato C, Chan CX, Ragan MA, Satoh N, Ball EE, Miller DJ
2018. ISME Journal • 12: 776–790. • DOI • PubMed • PDF • Feature
41
Selection of reference genes for transcript profiling of Sargassum polycystum by quantitative Real-Time Polymerase Chain Reaction
Sim MC, Chan CX, Ho CL, Phang SM
2018. Phycological Research • 66: 247–252. • DOI • PDF
2017
40
Signatures of adaptation and symbiosis in genomes and transcriptomes of Symbiodinium
González-Pech RA, Ragan MA, Chan CX
2017. Scientific Reports • 7: 15021. • DOI • PubMed • PDF
39
Biotic interactions as drivers of algal origin and evolution
Brodie J, Ball SG, Bouget FY, Chan CX, De Clerck O, Cock JM, Gachon C, Grossman AR, Mock T, Raven JA, Saha M, Smith AG, Vardi A, Yoon HS, Bhattacharya D
2017. New Phytologist • 216(3): 670-681 • DOI • PubMed • PDF
38
Insights into the red algae and eukaryotic evolution from the genome of Porphyra umbilicalis (Bangiophyceae, Rhodophyta)
Brawley SH, Blouin NA, Ficko-Blean E, Wheeler GL, Lohr M, Goodson HV, Jenkins JW, Blaby-Haas CE, Helliwell KE, Chan CX, Marriage TN, Bhattacharya D, Klein AS, Badis Y, Brodie J, Cao Y, Collén J, Dittami SM, Gachon CMM, Green BR, Karpowicz SJ, Kim JW, Kudahl UJ, Lin S, Michel G, Mittag M, Olson BJSC, Pangilinan JL, Peng Y, Qiu H, Shu S, Singer JT, Smith AG, Sprecher BN, Wagner V, Wang W, Wang ZY, Yan J, Yarish C, Zäuner-Riek S, Zhuang Y, Zou Y, Lindquist EA, Grimwood J, Barry KW, Rokhsar DS, Schmutz J, Stiller JW, Grossman AR, Prochnik SE
2017. Proceedings of the National Academy of Sciences USA • 114(31): E6361-E6370. • DOI • PubMed • PDF • Feature
37
The algal revolution
Brodie J, Chan CX, De Clerck O, Cock JM, Coelho SM, Gachon C, Grossman AR, Mock T, Raven JA, Smith AG, Yoon HS, Bhattacharya D
2017. Trends in Plant Science • 22(8): 726-738. • DOI • PubMed • Publisher • PDF
2016
36
Recapitulating phylogenies using k-mers: from trees to networks [version 2; referees: 2 approved]
Bernard G, Ragan MA, Chan CX
2016. F1000Research • 5: 2789. • DOI • PDF • Network
35
The transcriptomic response of the coral Acropora digitifera to a competent Symbiodinium strain: the symbiosome as an arrested early phagosome
Mohamed AR, Cumbo V, Harii S, Shinzato C, Chan CX, Ragan MA, Bourne DG, Willis BL, Ball EE, Satoh N, Miller DJ
2016. Molecular Ecology • 25(13): 3127–3141. • DOI • PubMed • PDF
34
Alignment-free microbial phylogenomics under scenarios of sequence divergence, genome rearrangement and lateral genetic transfer
Bernard G, Chan CX, Ragan MA
2016. Scientific Reports 6: 28970. • DOI • PubMed • PDF
33
PhySortR: a fast, flexible tool for sorting phylogenetic trees in R
Stephens TG, Bhattacharya D, Ragan MA, Chan CX
2016. PeerJ 4: e2038. • DOI • PubMed • PDF • Software
2015
32
The ReFuGe 2020 consortium – using ‘omics’ approaches to explore the adaptability and resilience of coral holobionts to environmental change
Voolstra CR, Miller DJ, Ragan MA, Hoffmann A, Hoegh-Guldberg O, Bourne D, Ball E, Ying H, Forêt S, Takahashi S, Weynberg KD, van Oppen MJ, Morrow K, Chan CX, Rosic N, Leggat W, Sprungala S, Imelfort M, Tyson GW, Kassahn K, Lundgren P, Beeden R, Ravasi T, Berumen M, Abel E, Fyffe T
2015. Frontiers in Marine Science 2: 68. • DOI • PDF
2014
31
Inferring phylogenies of evolving sequences without multiple sequence alignment
Chan CX, Bernard G, Poirion O, Hogan JM, Ragan MA
2014. Scientific Reports 4: 6504. • DOI • PubMed • PDF • Software
30
Molecular phylogenetics before sequences: oligonucleotide catalogs as k-mer spectra
Ragan MA, Bernard G, Chan CX
2014. RNA Biology 11(3): 176-185. • DOI • PubMed • PDF
29
A new species of Burkholderia isolated from sugarcane roots promotes plant growth
Paungfoo-Lonhienne C, Lonhienne T, Yeoh YK, Webb RI, Lakshmanan P, Chan CX, Lim PE, Ragan MA, Schmidt S, Hugenholtz P
2014. Microbial Biotechnology 7(2): 142-154. • DOI • PubMed • PDF
2013
28
Analysis of horizontal genetic transfer in red algae in the post-genomics age
Chan CX, Bhattacharya D
2013. Mobile Genetic Elements 3(6): e27669 • DOI • PubMed • PDF •
Online 2 Jan 2014
27
Foreign gene recruitment to the fatty acid biosynthesis pathway in diatoms
Chan CX, Baglivi FL, Jenkins CE, Bhattacharya D
2013. Mobile Genetic Elements 3(5): e27313 • DOI • PubMed • PDF •
Online 10 Dec 2013
26
Biological intuition in alignment-free methods: response to Posada
Ragan MA, Chan CX
2013. Journal of Molecular Evolution 77(1-2): 1-2 • DOI • PubMed • PDF •
25
Evidence for widespread exonic small RNAs in the glaucophyte alga Cyanophora paradoxa
Gross J, Wajid S, Price DC, Zelzion E, Li J, Chan CX, Bhattacharya D
2013. PLoS ONE 8(7): e67669 • DOI • PubMed • PDF •
24
Genome of the red alga Porphyridium purpureum
Bhattacharya D, Price DC, Chan CX, Qiu H, Rose N, Ball S, Weber APM, Arias MC, Henrissat B, Coutinho PM, Krishnan A, Zäuner S, Morath S, Hilliou F, Egizi A, Perrineau MM, Yoon HS
2013. Nature Communications 4: 1941 • DOI • PubMed • PDF •
ISI Highly Cited
23
Clustering evolving proteins into homologous families
Chan CX, Mahbob M, Ragan MA
2013. BMC Bioinformatics 14: 120 • DOI • PubMed • PDF •
22
Next-generation phylogenomics
Chan CX, Ragan MA
2013. Biology Direct 8: 3. • DOI • PubMed • PDF •
21
Phylogenomics of marine algae
Chan CX
2013. Malaysian Journal of Science 32 (SCS Special Issue): 11-18. • DOI • PDF
2012
20
Porphyra (Bangiophyceae) transcriptomes provide insights into red algal development and metabolism
Chan CX, Blouin NA, Zhuang Y, Zäuner S, Prochnik SE, Lindquist E, Lin S, Benning C, Lohr M, Yarish C, Gantt E, Grossman AR, Lu S, Müller K, Stiller J, Brawley SH, Bhattacharya D
2012. Journal of Phycology 48(6): 1328-1342 • DOI • PDF • Data Site •
19
Analysis of Alexandrium tamarense (Dinophyceae) genes reveals the complex evolutionary history of a microbial eukaryote
Chan CX, Soares MB, Bonaldo MF, Wisecaver JH, Hackett JD, Anderson DM, Erdner DL, Bhattacharya D
2012. Journal of Phycology 48(5): 1130-1142 • DOI • PubMed • PDF • Supp Info •
18
Major developmental regulators and their expression in two closely related species of Porphyra (Rhodophyta)
Stiller JW, Perry J, Rymarquis LA, Accerbi M, Green PJ, Prochnik S, Lindquist E, Chan CX, Yarish C, Lin S, Zhuang Y, Blouin NA, Brawley SH
2012. Journal of Phycology 48(4): 883-896 • DOI • PubMed • PDF •
17
Endosymbiotic and horizontal gene transfer in microbial eukaryotes: impacts on cell evolution and the tree of life
Chan CX, Bhattacharya D, Reyes-Prieto A
2012. Mobile Genetic Elements 2(2): 101-105 • DOI • PubMed • PDF •
16
Analysis of Porphyra membrane transporters demonstrates gene transfer among photosynthetic eukaryotes and numerous sodium-coupled transport systems
Chan CX, Zäuner S, Wheeler GL, Grossman AR, Prochnik SE, Blouin NA, Zhuang Y, Benning C, Berg GM, Yarish C, Eriksen RL, Klein AS, Lin S, Levine I, Brawley SH, Bhattacharya D
2012. Plant Physiology 158(4): 2001-2012 • DOI • PubMed • PDF • Supp Info
15
Cyanophora paradoxa genome elucidates origin of photosynthesis in algae and plants
Price DC, Chan CX, Yoon HS, Yang EC, Qiu H, Weber AP, Schwacke R, Gross J, Blouin NA, Lane C, Reyes-Prieto A, Durnford DG, Neilson JAD, Lang BF, Burger G, Steiner JM, Löffelhardt W, Meuser JE, Posewitz MC, Ball S, Arias MC, Henrissat B, Coutinho PM, Rensing SA, Symeonidi A, Doddapaneni H, Green BR, Rajah VD, Boore J, Bhattacharya D
2012. Science 335(6070): 843-847 • DOI • PubMed • PDF • Data Site •
2011
14
Red and green algal origin of diatom membrane transporters: insights into environmental adaptation and cell evolution
Chan CX, Reyes-Prieto A, Bhattacharya D
2011. PLoS ONE 6(12): e29138 • DOI • PubMed • PDF • Supp Info
13
Lateral transfer of genes and gene fragments in Staphylococcus extends beyond mobile elements
Chan CX, Beiko RG, Ragan MA
2011. Journal of Bacteriology 193(15): 3964-3977 • DOI • PubMed • PDF • Data Site •
12
Non-random sharing of Plantae genes
Chan CX, Bhattacharya D
2011. Communicative and Integrative Biology 4(3): 361-363 • DOI • PubMed • PDF
11
Plastid origin and evolution: new models provide insights into old problems
Chan CX, Gross J, Yoon HS, Bhattacharya D
2011. Plant Physiology 155(4): 1552-1560 • DOI • PubMed • PDF
10
Red and green algal monophyly and extensive gene sharing found in a rich repertoire of red algal genes
Chan CX, Yang EC, Banerjee T, Yoon HS, Martone PT, Estevez JM, Bhattacharya D
2011. Current Biology 21(4): 328-333 • DOI • PubMed • PDF • Supp Info
2009
9
Lateral transfer of genes and gene fragments in prokaryotes
Chan CX, Beiko RG, Darling AE, Ragan MA
2009. Genome Biology and Evolution 1:429-438 • DOI • PubMed • PDF
8
Are protein domains modules of lateral genetic transfer?
Chan CX, Darling AE, Beiko RG, Ragan MA
2009. PLoS ONE 4(2): e4524. • DOI • PubMed • PDF
2007
7
A two-phase strategy for detecting recombination in nucleotide sequences
Chan CX, Beiko RG, Ragan MA
2007. South African Computer Journal 38: 20-27 • link • PDF
2006
6
Detecting recombination in evolving nucleotide sequences
Chan CX, Beiko RG, Ragan MA
2006. BMC Bioinformatics 7:412 • DOI • PubMed • PDF
5
Trends in seaweed research
Chan CX, Ho CL, Phang SM
2006. Trends in Plant Science 11(4): 165-166 • DOI • PubMed • PDF
2005
4
A word-oriented approach to alignment validation
Beiko RG, Chan CX, Ragan MA
2005. Bioinformatics 21(10): 2230-2239 • DOI • PubMed • PDF • Software
3
Seaweed diversity of the Langkawi Islands with emphasis on the northeastern region
Phang SM, Wong CL, Lim PE, Yeong HY, Chan CX
2005. Malaysian Journal of Science 24 (Langkawi Special Issue): 77-94 • link • PDF
2004
2
Comparing different approaches in large-scale seaweed gene discovery
Chan CX
2004. Bioinformatics India 2(3): 106-111 • PDF
1
Optimisation of RNA extraction from Gracilaria changii (Gracilariales, Rhodophyta)
Chan CX, Teo SS, Ho CL, Othman RY, Phang SM
2004. Journal of Applied Phycology 16(4): 297-301 • DOI • PDF